Visualising Numbers Needed for Change
Source:R/convert.threshold.to.er.R, R/erDataSeq.R, R/ggNNC.R
nncvis.RdThese functions can be used to visualise Numbers Needed for Change (or
Numbers Needed to Treat).
erDataSeq is a helper function to generate an Event Rate Data
Sequence, and it uses convert.threshold.to.er and
convert.er.to.threshold to convert thresholds to event rates and vice
versa.
convert.threshold.to.er(
threshold,
mean,
sd,
eventIfHigher = TRUE,
pdist = stats::pnorm
)
convert.er.to.threshold(
er,
mean,
sd,
eventIfHigher = TRUE,
qdist = stats::qnorm
)
erDataSeq(
er = NULL,
threshold = NULL,
mean = NULL,
sd = NULL,
eventIfHigher = TRUE,
pRange = c(1e-06, 0.99999),
xStep = 0.01
)
ggNNC(
cerDataSeq,
d = NULL,
eventDesirable = TRUE,
r = 1,
xlab = "Continuous outcome",
plotTitle = c("Numbers Needed for Change = ", ""),
theme = ggplot2::theme_bw(),
lineSize = 1,
cerColor = "#EBF2F8",
eerColor = "#172F47",
cerLineColor = "#888888",
eerLineColor = "#000000",
dArrowColor = "#000000",
cerAlpha = 0.66,
eerAlpha = 0.66,
xLim = NULL,
xLimAutoDensityTolerance = 0.001,
showLegend = TRUE,
verticalLineColor = "#172F47",
desirableColor = "#00FF00",
desirableAlpha = 0.2,
undesirableColor = "#FF0000",
undesirableAlpha = 0.2,
desirableTextColor = "#009900",
undesirableTextColor = "#990000",
dArrowDistance = 0.04 * max(cerDataSeq$density),
dLabelDistance = 0.08 * max(cerDataSeq$density)
)Arguments
- threshold
If the event rate is not available, a threshold value can be specified instead, which is then used in conjunction with the mean (
mean) and standard deviation (sd) and assuming a normal distribution to compute the event rate.- mean
The mean of the control group distribution.
- sd
The standard deviation (of the control distribution, but assumed to be the same for both distributions).
- eventIfHigher
Whether scores above or below the threshold are considered 'an event'.
- pdist, qdist
Distributions to use when converting thresholds to event rates and vice versa; defaults to the normal distribution.
- er
Event rate to visualise (or convert).
- pRange
The range of probabilities for which to so the distribution.
- xStep
Precision of the drawn distribution; higher values mean lower precision/granularity/resolution.
- cerDataSeq
The
cerDataSeqobject.- d
The value of Cohen's d.
- eventDesirable
Whether an event is desirable or undesirable.
- r
The correlation between the determinant and behavior (for mediated NNC's).
- xlab
The label to display for the X axis.
- plotTitle
The title of the plot; either one character value, this value if used; if two, they are considered a prefix and suffix to be pre/appended to the NNC value.
- theme
The theme to use for the plot.
- lineSize
The thickness of the lines in the plot.
- cerColor
The color to use for the event rate portion of the control group distribution.
- eerColor
The color to use for the event rate portion of the experimental group distribution.
- cerLineColor
The line color to use for the control group distribution.
- eerLineColor
The line color to use for the experimental group distribution.
- dArrowColor
The color of the arrow to show the effect size.
- cerAlpha
The alpha value (transparency) to use for the control group distribution.
- eerAlpha
The alpha value (transparency) to use for the control group distribution.
- xLim
This can be used to manually specify the limits for the X axis; if
NULL, sensible limits will be derived usingxLimAutoDensityTolerance.- xLimAutoDensityTolerance
If
xLimisNULL, the limits will be set where the density falls below this proportion of its maximum value.- showLegend
Whether to show the legend (only if showing two distributions).
- verticalLineColor
The color of the vertical line used to indicate the threshold.
- desirableColor
The color for the desirable portion of the X axis.
- desirableAlpha
The alpha for the desirable portion of the X axis.
- undesirableColor
The color for the undesirable portion of the X axis.
- undesirableAlpha
The color for the undesirable portion of the X axis.
- desirableTextColor
The color for the text to indicate the desirable portion of the X axis.
- undesirableTextColor
The color for the text to indicate the undesirable portion of the X axis.
- dArrowDistance
The distance of the effect size arrow from the top of the distributions.
- dLabelDistance
The distance of the effect size label from the top of the distributions.
Value
erDataSeq returns a data sequence; ggNNC a
ggplot2::ggplot().
Details
These functions are used by nnc() to show the distributions, and
event rates. They probably won't be used much on their own.
References
Gruijters, S. L., & Peters, G. Y. (2019). Gauging the impact of behavior change interventions: A tutorial on the Numbers Needed to Treat. PsyArXiv. doi: 10.31234/osf.io/2bau7
See also
Examples
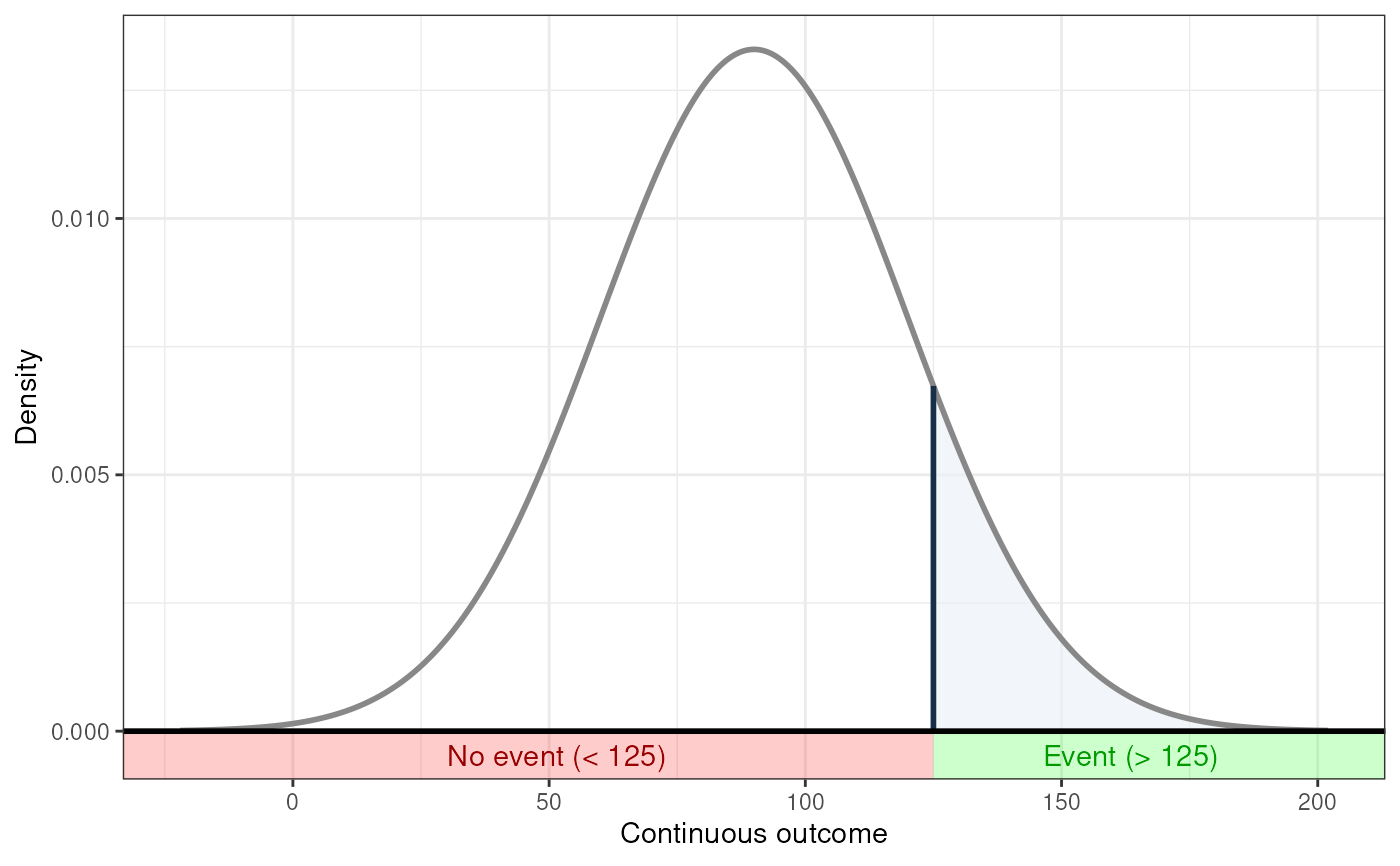
### Show distribution for an event rate value of 125
behaviorchange::ggNNC(behaviorchange::erDataSeq(threshold=125, mean=90, sd=30));
 ### If the event occurs under the threshold instead of
### above it
behaviorchange::ggNNC(behaviorchange::erDataSeq(threshold=125,
mean=90, sd=30,
eventIfHigher = FALSE));
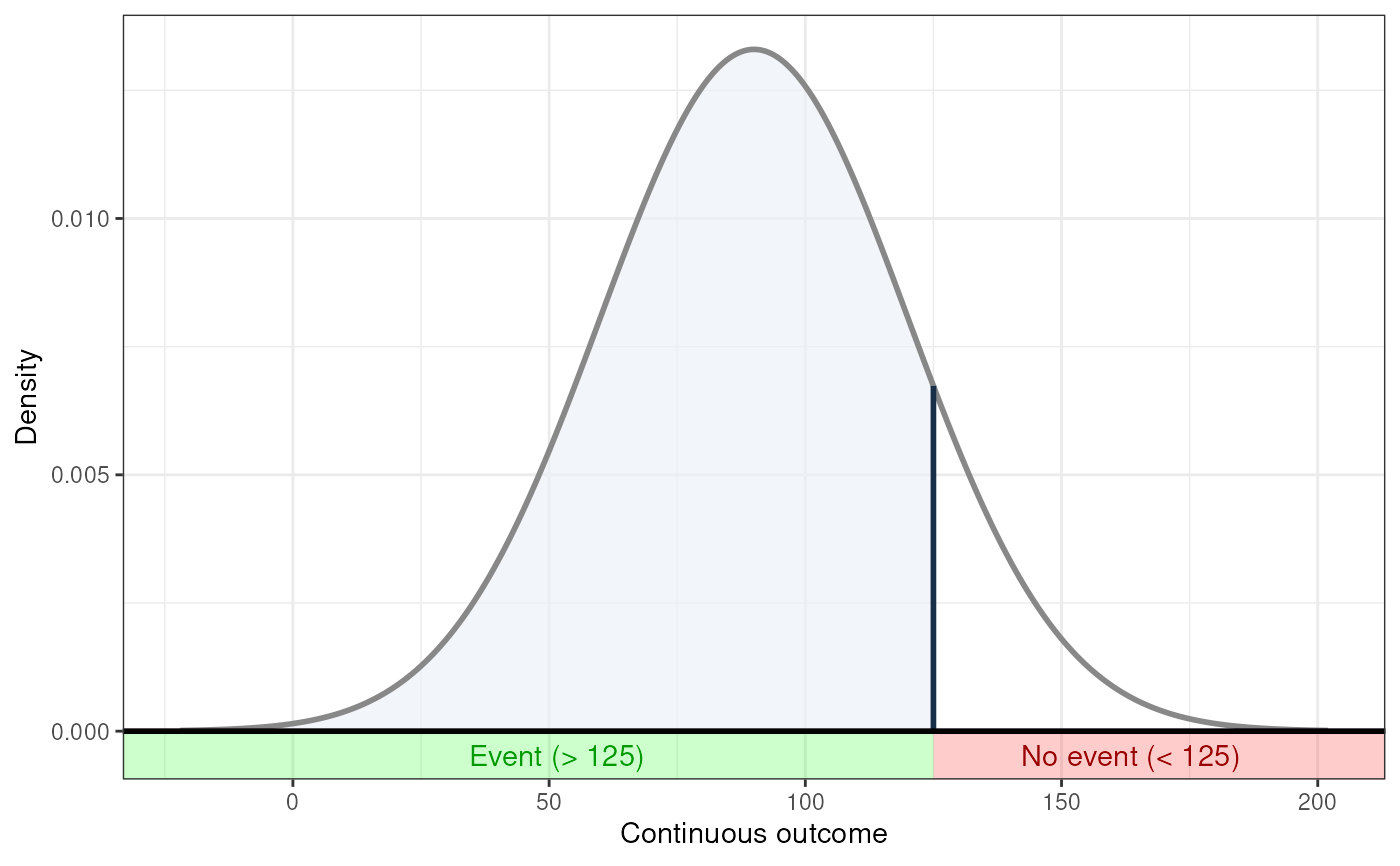
### If the event occurs under the threshold instead of
### above it
behaviorchange::ggNNC(behaviorchange::erDataSeq(threshold=125,
mean=90, sd=30,
eventIfHigher = FALSE));
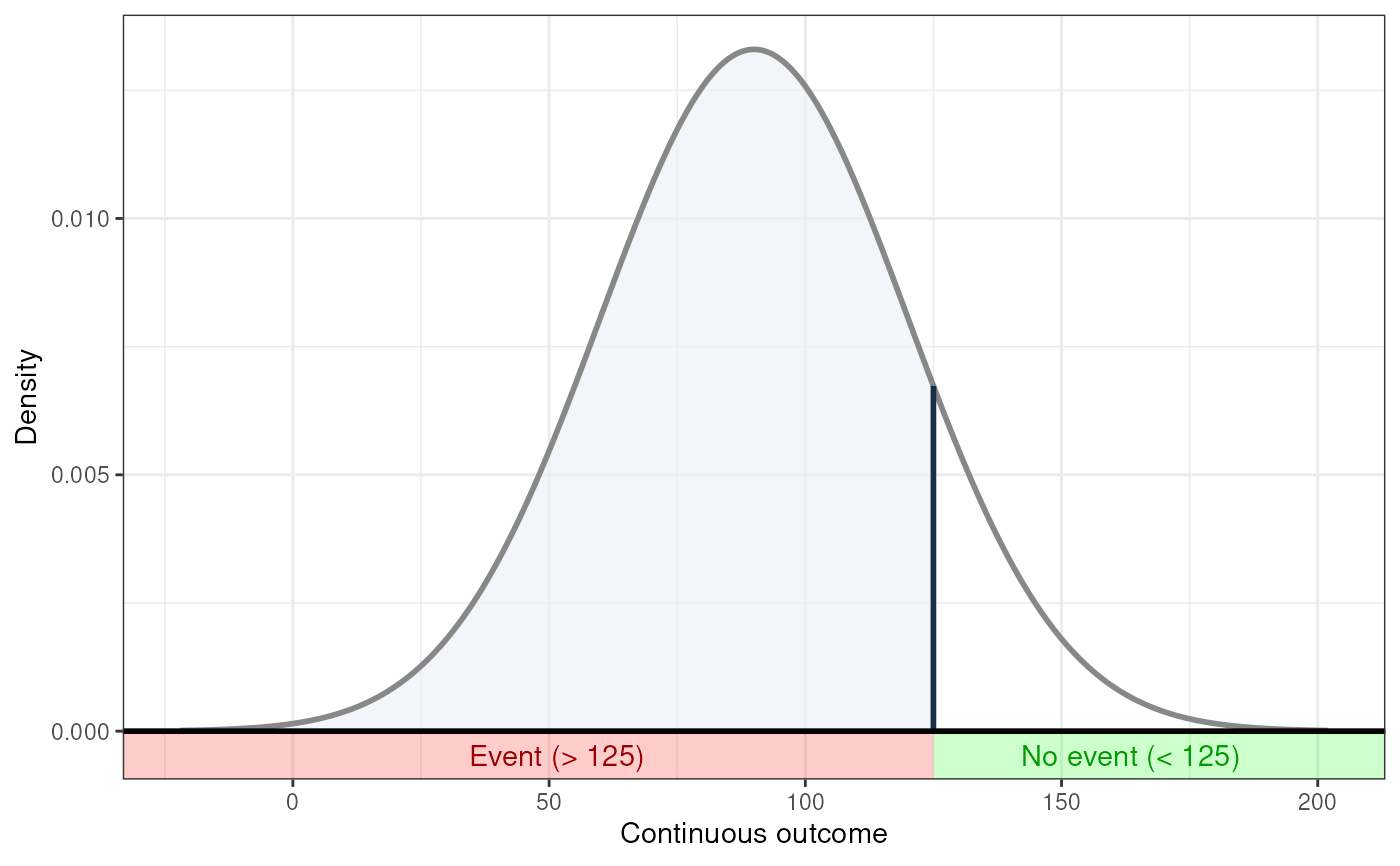
 ### ... And for undesirable events (note how
### desirability is an argument for ggNNC, whereas
### whether an event occurs 'above' or 'below' the
### threshold is an argument for erDataSeq):
behaviorchange::ggNNC(behaviorchange::erDataSeq(threshold=125,
mean=90, sd=30,
eventIfHigher = FALSE),
eventDesirable = FALSE);
### ... And for undesirable events (note how
### desirability is an argument for ggNNC, whereas
### whether an event occurs 'above' or 'below' the
### threshold is an argument for erDataSeq):
behaviorchange::ggNNC(behaviorchange::erDataSeq(threshold=125,
mean=90, sd=30,
eventIfHigher = FALSE),
eventDesirable = FALSE);
 ### Show event rate for both experimental and
### control conditions, and show the numbers
### needed for change
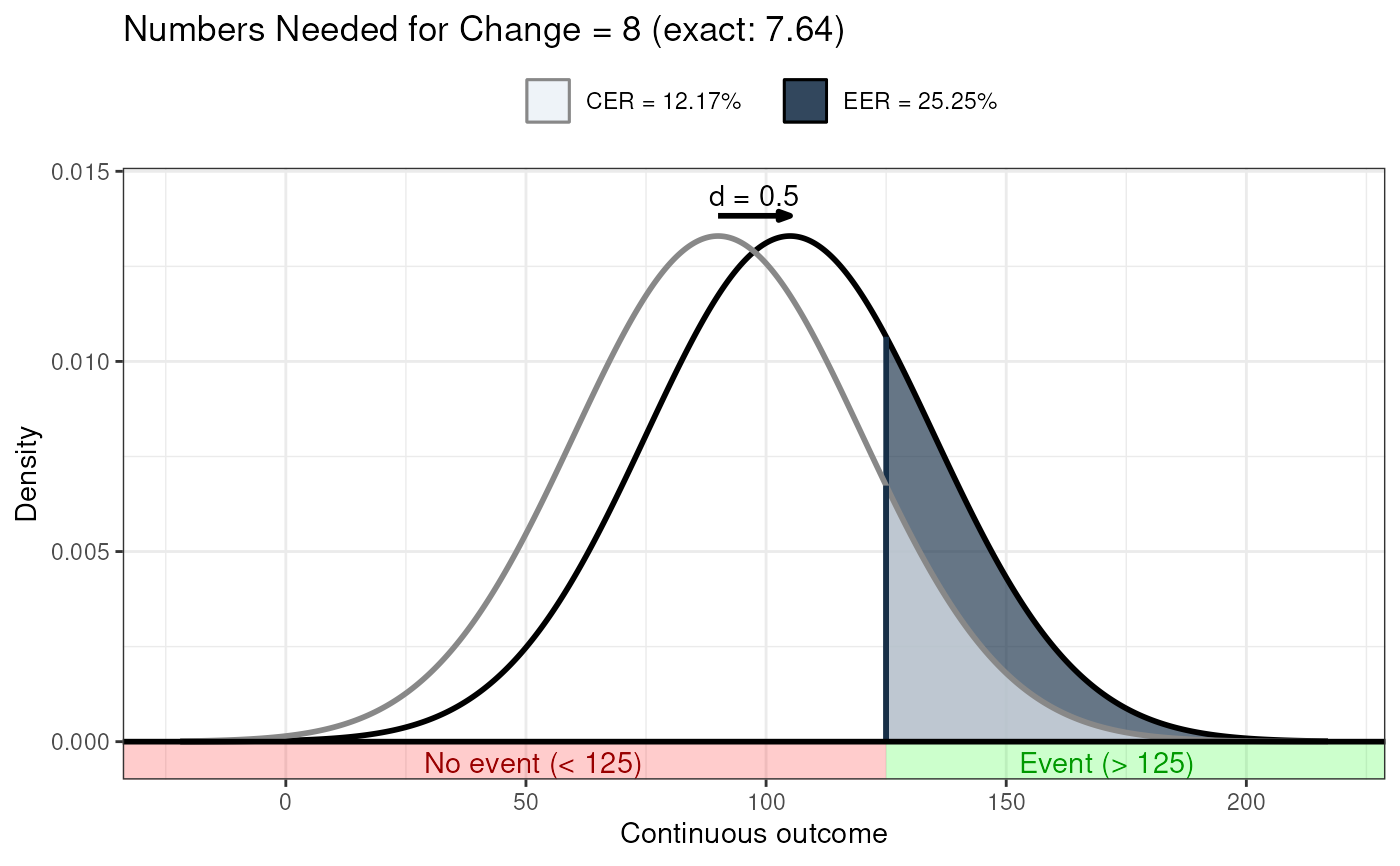
behaviorchange::ggNNC(behaviorchange::erDataSeq(threshold=125,
mean=90, sd=30),
d=.5);
### Show event rate for both experimental and
### control conditions, and show the numbers
### needed for change
behaviorchange::ggNNC(behaviorchange::erDataSeq(threshold=125,
mean=90, sd=30),
d=.5);
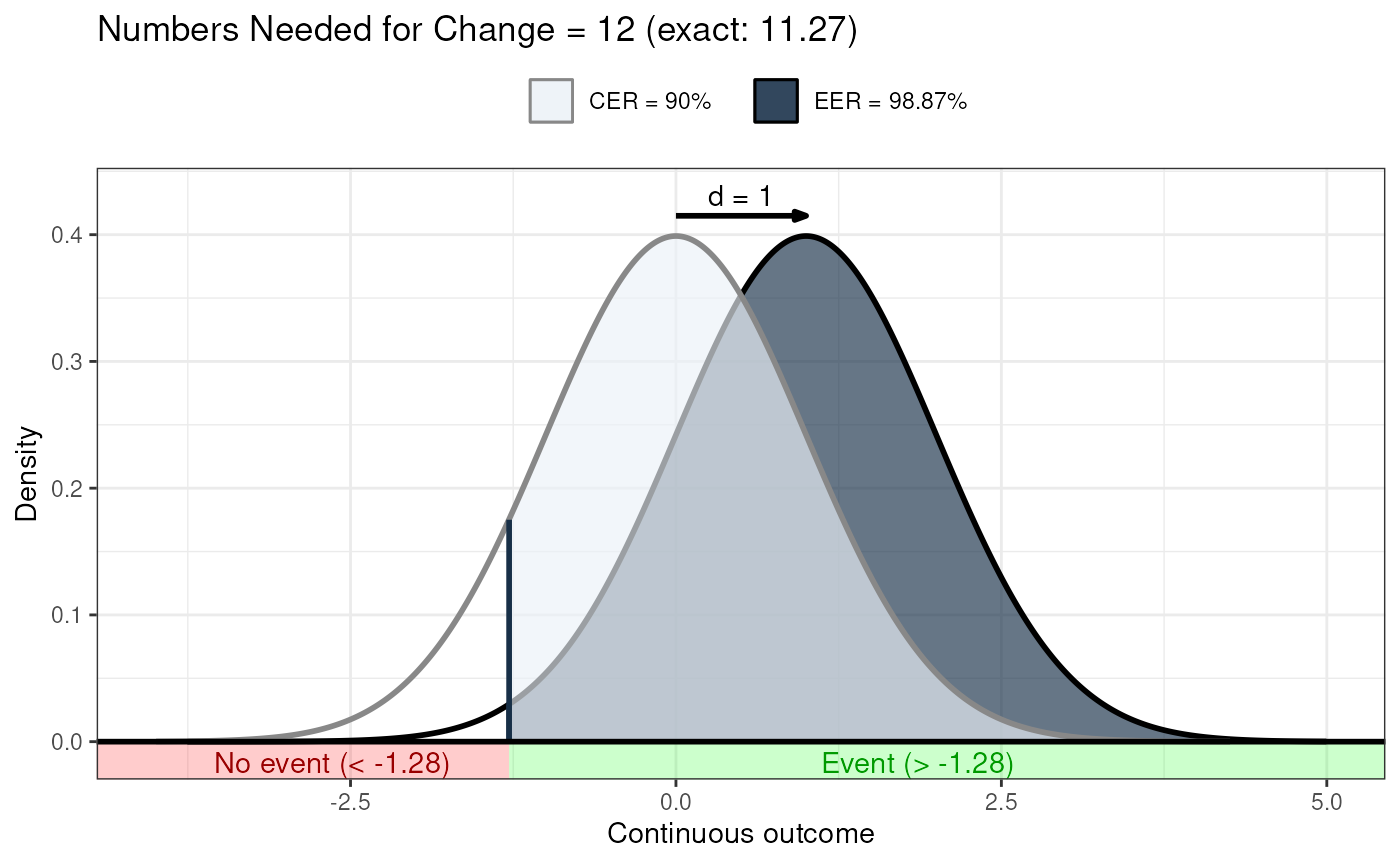
 ### Illustration of how even with very large effect
### sizes, if the control event rate is very high,
### you'll still need a high number of NNC
behaviorchange::ggNNC(behaviorchange::erDataSeq(er=.9),
d=1);
### Illustration of how even with very large effect
### sizes, if the control event rate is very high,
### you'll still need a high number of NNC
behaviorchange::ggNNC(behaviorchange::erDataSeq(er=.9),
d=1);